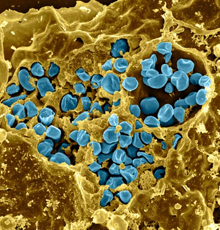
F. novicida and F. tularensis (blue in this electron microscopy image) are bacteria that live inside mammalian cells.
In F. novicida, a genetic defense system used by other bacteria to defend themselves against phages has acquired a distinct function not seen before: evading the host's immune system.
CRISPR, a system of genes that bacteria use to defend themselves against viruses, has been found to be involved in helping some bacteria evade the mammalian immune system.
The results were published online Sunday, April 14 ahead of print in Nature.
CRISPR is itself a sort of immune system for bacteria. Its function was discovered by dairy industry researchers seeking to prevent phages, the viruses that infect bacteria, from ruining the cultures used to make cheese and yogurt. Bacteria incorporate small bits of DNA from phages into their CRISPR region and use that information to fight off the phages by chewing up their DNA.
Now scientists at the Division of Infectious Diseases of the Emory University School of Medicine and the Emory Vaccine Center have shown that Francisella novicida, a close relative of the bacterium that causes tularemia, and another bacterium that causes meningitis, need parts of the CRISPR system to stay infectious. F. novicida, which grows inside mammalian cells, employs parts of CRISPR to shut off a bacterial gene that would otherwise trigger detection and destruction of the bacteria by its host.
Because disabling CRISPR creates a weakened bacterial strain that is easily recognized by the immune system, the finding could accelerate vaccine development. But it is also a broader reminder that in biology, defensive tools can be co-opted for purposes of stealth.
"CRISPR systems are bacterial defenses, but we've found that bacteria can use them offensively to hide from the host immune system and cause disease," says David Weiss, PhD, assistant professor of medicine (infectious diseases) at Emory University School of Medicine and Yerkes National Primate Research Center.
The CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) system has attracted recent attention among scientists for its potential uses in genetic engineering and biotechnology, but its roles in gene regulation and evading host immunity have remained relatively unexplored, Weiss says.
Weiss first isolated strains of F. novicida that had defects in their CRISPR systems while working as a postdoc with Denise Monack at Stanford. F. novicida infects rodents and only rarely infects humans. It is a model for studying the more dangerous F. tularensis, a potential biological weapon. Weiss was looking for F. novicida genes that are important for virulence: causing disease in a live animal.
Intriguingly, he found a DNA sequence that has recently been shown to encode a protein of the CRISPR system. What they were doing in F. novicida during infection was a puzzle.
"The mutations have a strong effect in the bacteria," Weiss says. "The wild type will kill mice, while the mutants are eradicated after a couple days. But why would the bacteria need to defend against foreign DNA to cause disease in a mouse? It didn't make sense."
The researchers discovered that the bacteria require one of the CRISPR genes to turn off production of a lipoprotein – part of the bacterial cell membrane -- when the bacteria infect mammalian cells. For immune cells, lipoprotein is like blood in the water for a shark. A little whiff excites them. So for the bacteria to survive undetected, they have to silence lipoprotein production.
Working with Weiss, graduate student Tim Sampson – who is first author of the Nature paper -dissected which parts of the CRISPR system were needed to turn off the lipoprotein. The CRISPR system consists of genes encoding several proteins and also incorporates small bits of DNA from phages as "repeats" into the bacterial DNA. RNA produced from the repeats guides an enzyme called Cas9 to slice up the phage DNA.
Sampson and Weiss found that part of the F. novicida CRISPR system makes an RNA that directs Cas9 against the lipoprotein gene. Weiss says the Cas9 regulatory system allows F. novicida to tune down the lipoprotein efficiently at the times when detection could be harmful, while still keeping it around for its function – still unclear – when the bacteria are outside the host.
"The finding that Cas9 is regulating a bacterial gene rather than slicing up a phage gene appears to be new, although there were already some hints that CRISPRs had broader functions in other bacteria," Sampson says.
To show that their results were not peculiar to F. novicida, the researchers collaborated with the laboratory of Yih-Ling Tzeng, PhD, assistant professor of medicine (infectious diseases) at Emory. They generated a strain of Neisseria meningitidis -- a cause of meningitis infections in humans -- with a deletion in Cas9. The mutated strain displayed defects in its ability to adhere to, invade and replicate in human cells. This suggests that similar functions for Cas9 and CRISPR may be found in other bacteria.
"Most of the bacteria that encode Cas9 are either pathogenic, or can commonly be found in the human body," Sampson says. "I think our findings will encourage other scientists to re-examine the functions of Cas9 and CRISPR in other bacteria, to look at interactions with the host."
For example, some Streptococcus bacteria and Listeria have similar CRISPR systems, but any potential function in causing disease in humans has not been revealed. Weiss and Sampson plan to investigate further how Cas9 functions to shut off the lipoprotein gene in F. novicida and how Cas9 becomes activated.
Sampson is a student in Emory's Microbiology and Molecular Genetics graduate program. The research was supported by the Southeastern Regional Center of Excellence for Emerging Infections and Biodefense and National Institute of Allergy and Infectious Diseases (U54-AI057157, R56-AI061031, R56-AI87673).
Reference: T.R. Sampson, S.D. Saroj, A.C. Llewellyn, Y.L. Tzeng, and D.S. Weiss. A CRISPR-CAS system mediates bacterial innate immune evasion and virulence. Nature advanced online publication (2013).
